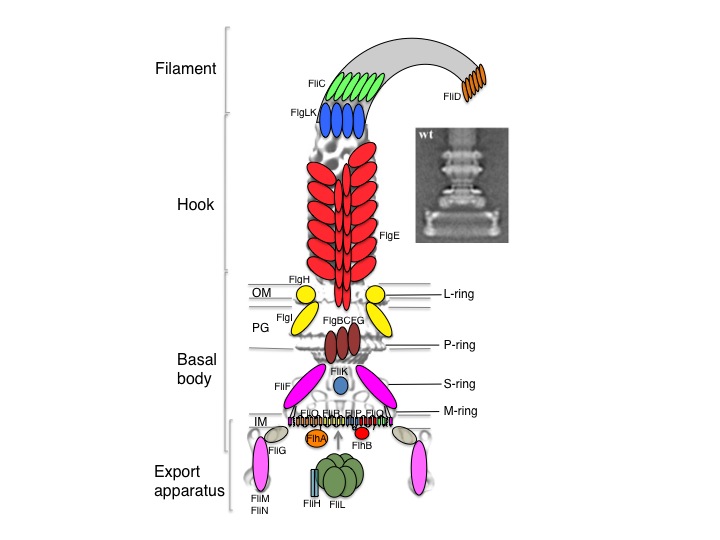
The bacterial flagellum is a large macromolecular nano-machine present at the surface of many bacteria. Its role is to allow bacterial motility, through the rotation of a long filamentous structure that protrudes away from the bacterial surface. Importantly, the flagellum is an essential pathogenicity factor in most bacterial pathogens, including E.coli, Salmonella, Pseudomonas, Helicobacter, Vibrio, Campylobacter etc…. As a consequence, understanding the flagellum at the molecular level could provide many prospects for new antibacterial therapeutics. Conversely, a number of groups have successfully exploited the flagellum-driven motility to design bacterial-based delivery systems. In particular, the anticancer potential of Salmonella typhimurium, which is flagellum dependent, has been extensively studied. For these reasons, understanding the flagellum at the molecular level is of primary importance. Yet the complexity and membrane embedding of these structures renders them very challenging for structural studies.
Our aim is to use a combination of Cryo-EM, X-ray crystallography, NMR and other biophysical and biochemical methods, to piece together the structure of the flagellum, understand its mechansim of action, and deceipher how it assembles.